BioWish: a molecular biology command extension to Tcl/Tk
Thomas Sicheritz-Pontén  Dep. of Molecular Biology, University of Uppsala, Sweden
thomas@evolution.bmc.uu.se
Red text indicates corrections or updates
Dep. of Molecular Biology, University of Uppsala, Sweden
thomas@evolution.bmc.uu.se
Red text indicates corrections or updates
Introduction
The Tcl/Tk [Ousterhout,
1994] scripting language has proved to be a powerful tool for building
programs involved in the analysis of molecular sequence data. However,
typical ``biological'' operations like the translation of a nucleotide
sequence to the corresponding amino acid sequence, or the calculation of
the G+C content in different codon positions in a 50 kbp cosmid sequence
are performed far too slowly with the standard Tcl commands. To circumvent
this problem, we have constructed a library that extends the Tcl/Tk language
by adding primitive operators suited for sequence analysis implemented
in the C-language. Additional commands related to molecular biology, written
in Tcl are included. Built as a shared library the usage is easy and does
not require modification of the Tcl/Tk source code.
Availability
BioWish can be obtained from the WWW site http://evolution.bmc.uu.se/~thomas/mol_linux
The distribution consists of a single C-source file which should compile
without modifications on all Unix systems capable of dynamical loading.
It requires Tcl 7.5/Tk4.1 Tcl8.0/Tk8.0
or higher. No patching of the Tcl/Tk core is required. On systems where
dynamic loading is not available, Biowish can be compiled as a standalone
Tk intepreter.
BioWish Features
-
sequence editing: reverse, complement, antiparallel
-
translations
-
sequence statistics
-
G+C content in different positions
-
dna incrementor
-
sequence mutation
-
database searches with BLAST
-
sequence editing widget
-
text editing widget
-
graphical open reading frame widget
-
sequence conversion via buildin readseq
-
contig projects
Usage
Using BioWish in Tcl scripts requires only a single additional line, a
directive to the Tcl intepreter to load the command extension from the
shared library, via the load command.
load ./biowish.so
bio_readfasta ecoli.fas seq
set s [string tolower $seq(sequence)]
puts [bio_seqinfo $s]
for { set nt aaa} {$nt!="aaaaaaa"} {bio_dna_incr nt} {
puts "$nt = [regsub -all $nt $s {} temp]"
}
load ./biowish.so ;# package require Biowish
biowish::readseq ecoli.fas seq
set s [string tolower $seq(sequence)]
puts [biowish::seqinfo $s]
for { set nt aaa} {$nt!="aaaaaaa"} {biowish::dna_incr nt} {
puts "$nt = [regsub -all $nt $s {} temp]"
}
BioWish also includes a sequence editing Tk widget which takes advantage
of the extended Tcl commands. (Figure 1).
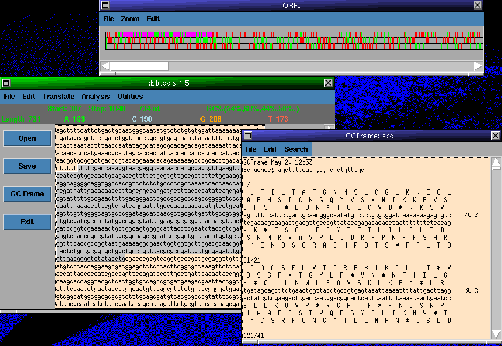
Figure 1: A screen shot of the Tk sequence editing widget
References
-
Ousterhout, 1994
-
Ousterhout, J. K. (1994). Tcl and the Tk Toolkit. Addison-Wesley,
Reading, MA, USA.
About this document ...
BioWish: a molecular biology command extension to Tcl/Tk
This document was generated using the LaTeX2HTML
translator Version 96.1 (Feb 5, 1996) Copyright © 1993, 1994, 1995,
1996, Nikos Drakos,
Computer Based Learning Unit, University of Leeds.
The command line arguments were:
latex2html -split 0 application_note.tex.
The translation was initiated by Thomas Sicheritz on Tue Jun 10 09:47:07
MET DST 1997
-
...Sicheritz-Pontén
-
This work was supported by contract no. BIO4 CT-95-0130 from the European
Commission to C. Kurland.
Thomas Sicheritz
Tue Jun 10 09:47:07 MET DST 1997


