The ghemical homepage
Introduction
Ghemical is computational chemistry package, which is licensed under
GNU GPL
. The ghemical authors are listed
here
.
Ghemical is implemented using the C++ programming language, and it has
a graphical user interface which utilizes the
OpenGL
graphics interface and the
GTK+
multiplatform widget library. Here are some features of the program:
- Compute energy and gradient using :
- quantum mechanical methods
- Ab Initio methods provided by MPQC.
- Semi-Empirical methods provided by MOPAC7.
- molecular mechanics methods
- all-atom MM methods (Tripos 5.2 + others).
- coarse-grained MM for protein molecules [1] (work in progress).
- optional distance constraints can be applied in MM calculations.
- Options for studying a molecular system using :
- Geometry Optimization.
- Molecular Dynamics.
- Conformational Searching :
- Random Search.
- Systematic Search.
- Monte Carlo Search.
- Other options for QM methods :
- Stationary State Search.
- Transition State Search.
- Population Analysis.
- Interfaces to external programs :
- Many file import/export features provided by OpenBabel.
- OpenGL graphics presentations :
- molecular graphics presentations :
- Ball-And-Stick.
- Van der Waals.
- Cylinders.
- Wireframe.
- molecular editing tools :
- add and remove atoms and bonds.
- add or remove Hydrogens automatically.
- sequence builder and identifier :
- protein sequences.
- nucleic acid sequences.
- measurement tools for measuring :
- distances.
- angles.
- torsions.
- visualization options :
- colored planes.
- colored surfaces.
- volume rendering.
- protein ribbon models.
- interactive graphing options :
- Energy vs. torsion plot.
- Energy vs. 2 torsions plot.
- energy level diagram plot.
- reaction coordinate plot.
- "Project View" for viewing/editing the model and related information.
- Text window for text-formatted output and logging, and a text command parser/interpreter.
- Molecular Weight calculator.
- MD Trajectory file viewer.
Screenshots
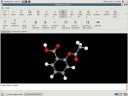 |
The graphical user interface, and an example molecule (acetylsalicylic acid). |
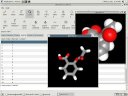 |
Different views and graphics rendering options demonstrated. |
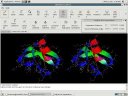 |
Protein ribbon models and a stereo display feature demonstrated. |
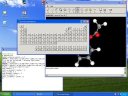 |
Since the GTK+ environment can be ported to several operating systems, the program is portable as well
(the screenshot is from a Windows/Cygwin system). |
Download
You can get the packages from the
download
page.
The current stable version 2.00 was released 2006-04-xx.
It is tested to work best with the following other packages/programs:
Development
Ghemical source code is developed at a
CVS server
hosted by the
bioinformatics.org
site.
Discussions about ghemical development can be followed at
ghemical-devel
mailing list.
Links
References
[1] Hassinen, T.; Peräkylä, M.
J Comput Chem 2001, 22, 1229-1242
This page was last modified 2006-04-12.



