ABE
Homepage
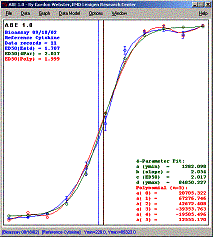
ABE is a small, fast and
convenient program for visualizing and modeling experimental bioassay data. The
data can be modeled using either polynomials or a more specific four-parameter model
based upon the standard, sigmoidal dose-response curve.
 ABE (Copyright ã 2002, 2003 Gordon Webster, EMD Lexigen Research
Center) is Open Source software that is available at no charge. ABE is supplied
as is, without warranty, and may be freely used and distributed under
the terms of the Gnu General
Public License (GPL) as described in Appendix A of the ABE manual.
ABE (Copyright ã 2002, 2003 Gordon Webster, EMD Lexigen Research
Center) is Open Source software that is available at no charge. ABE is supplied
as is, without warranty, and may be freely used and distributed under
the terms of the Gnu General
Public License (GPL) as described in Appendix A of the ABE manual.
The current version of ABE
(1.0) is available as Python source code (any platform supporting release 2.2
(or later) of Python) or as a convenient, precompiled binary (Win NT/2000).
Source code, binaries and
documentation are available at the ABE
project pages at Bioinformatics.org
A Python runtime environment must be installed if you
plan to work with the ABE source code. Downloads of the current version of
Python and detailed documentation, are available at the Python Website.
Gordon Webster can be contacted via his
bioinformatics.org email: gwebster@bioinformatics.org

